
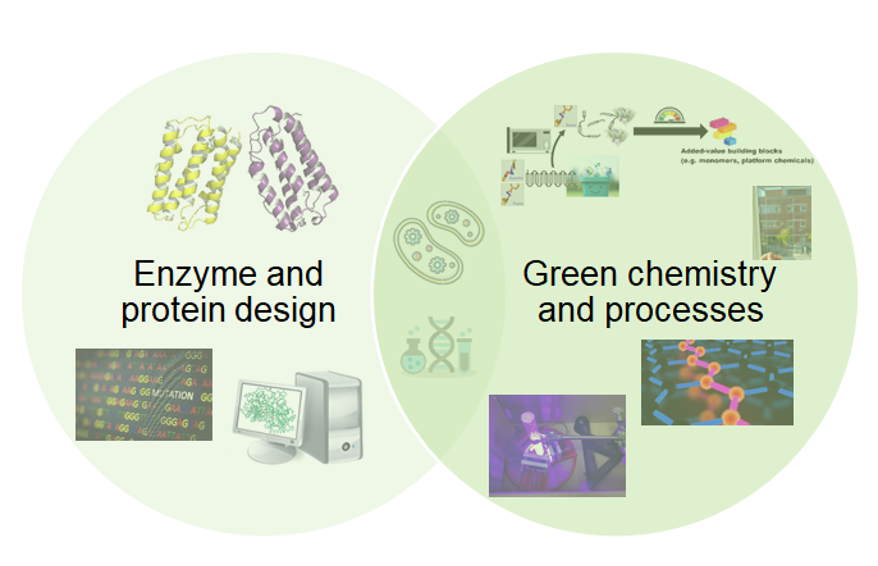
At the Syren Lab at KTH Royal Institute of Technology, we explore fundamental enzyme catalysis applied to organic chemical synthesis. Our research lies at the intersection of protein engineering, chemistry, material science and catalysis. By combining computational protein design, structural biology, and mechanistic enzymology with experimental biochemistry, we investigate how enzyme structure and dynamics control reactivity, selectivity, and stability of biocatalysts. A central goal is to uncover fundamental structure–function relationships that allow us to tailor enzymes for promiscuous reactions and new-to-nature chemistries.
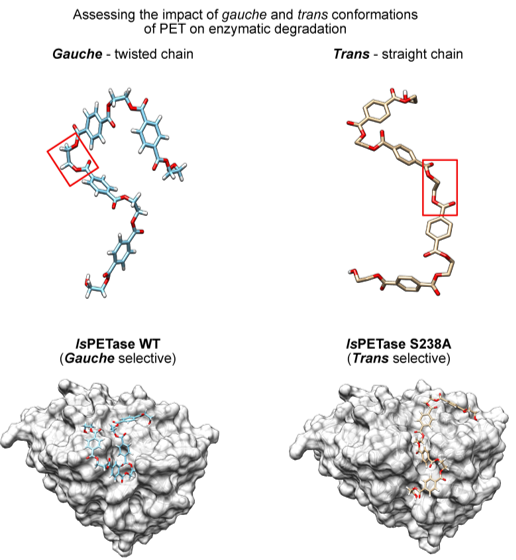
A major focus of our work is the development of biocatalytic strategies for sustainable organic chemistry and polymer science. We design and engineer enzymes capable of making synthetic and bio-based polymers, as well as recycling them at end-of-life; contributing to circular materials solutions and greener chemical processes. Through an integrated approach that spans molecular design to materials characterization, we aim to build robust enzymatic platforms that support sustainable production of building blocks, selective recycling under mild conditions, and upcycling of polymeric materials.
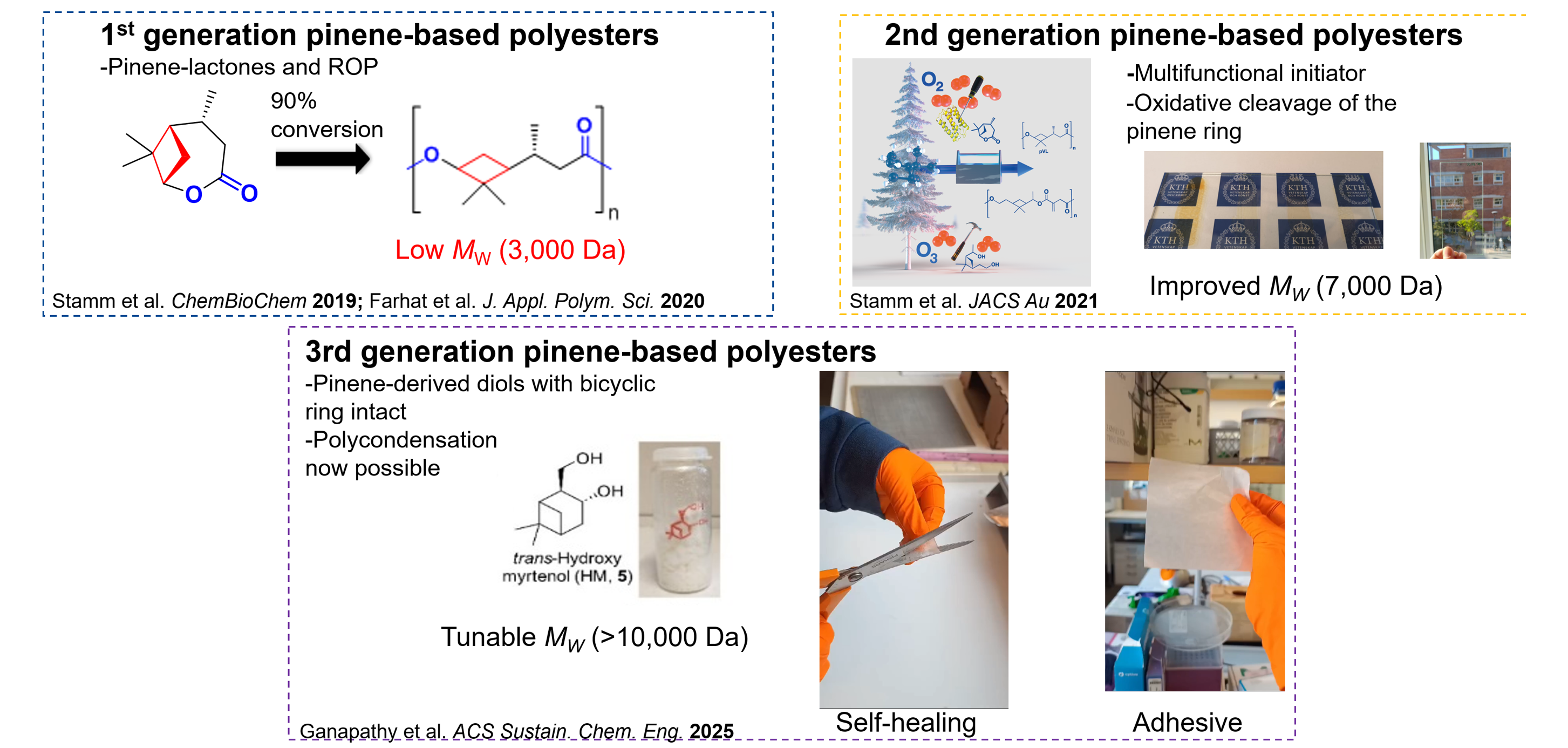
To achieve this, we work with in vitro metabolic engineering to design artificial biosynthetic pathways for the upcycling of abundant natural terpenes into added-value building blocks. We combine biochemical process engineering, enzyme design and polymer technology to afford green routes towards renewable terpene-based materials. We are using different polymerization techniques, including controlled radical polymerization and ring opening polymerization to afford new bio-based materials such as bio-based polyesters with properties approaching aromatic fossil-based plastics. We further analyse the thermal properties, molecular weight distributions and the molecular structures of the obtained materials.
Our funding
The Swedish Foundation for Strategic Environmental Research (Mistra; project Mistra SafeChem, Project No. 2018/11)
the Swedish Research Council (VR)
the NovoNordisk Foundation